Pept-ins™
Short amyloidogenic protein fragments for the selective suppression or detection of proteins
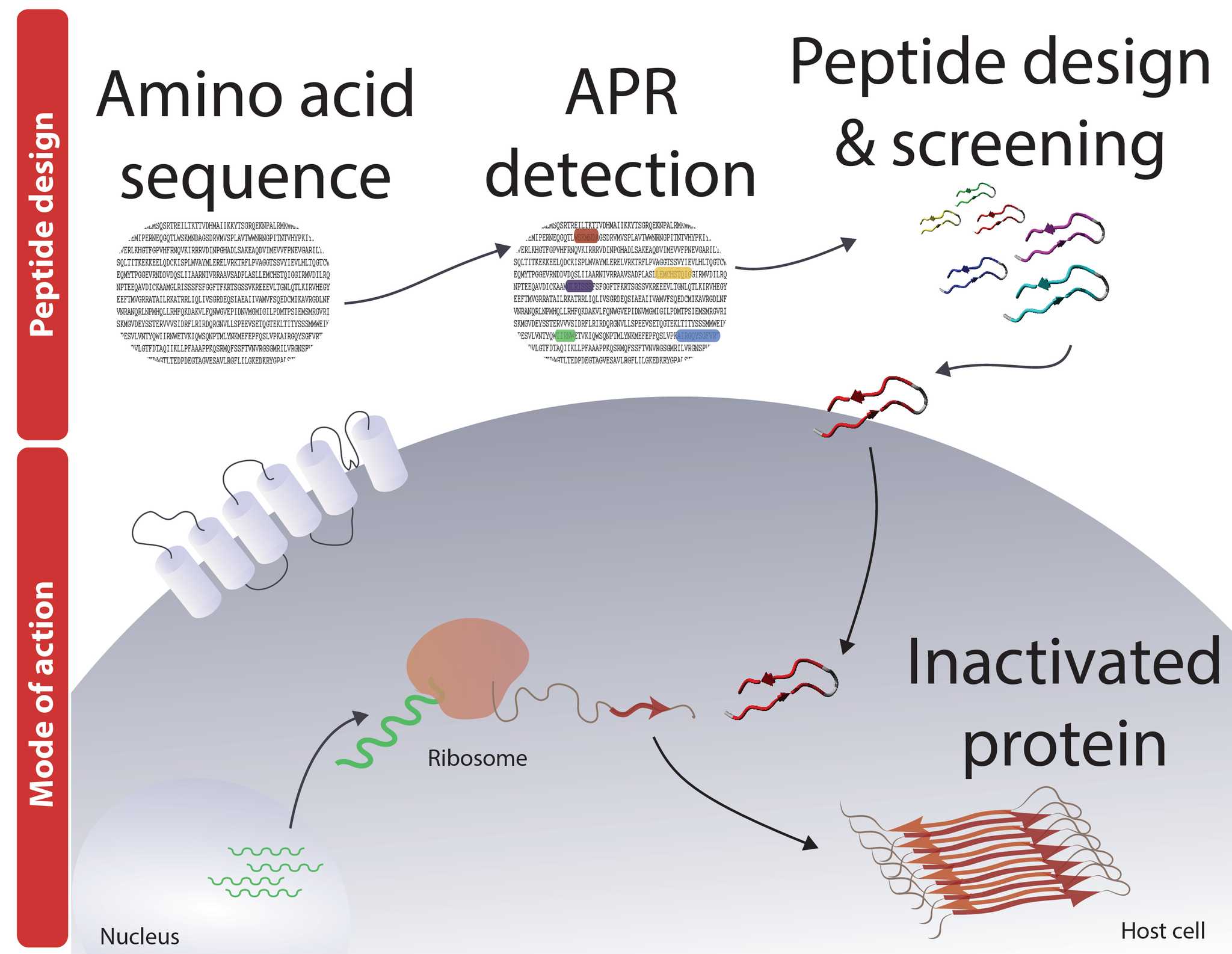
The Pept-ins™ technology developed by Frederic Rousseau and Joost Schymkowitz is the application of targeted protein aggregation as a radically new method to design novel biotherapeutics. Pept-ins™ (from peptide interferors) are synthetic amyloidogenic peptides that induce aggregation of their target proteins. Pept-ins™ can deactivate the targeted protein and destabilize the targeted pathogen or targeted cell.
The Switch laboratory pioneered the development of several bioinformatics algorithms, including TANGO [1] and WALTZ [2] for the prediction of aggregation prone regions within the primary sequence of proteins, and this is where the workflow starts. The designed Pept-ins™ are then screened for their ability to induce aggregation of the target protein first in vitro and then in vivo.
The Switch laboratory has documented that Pept-ins™ are effective in plants [3,4], mammalian cells [5], Gram-positive and Gram-negative bacteria [6,7], as well as for viruses [8].
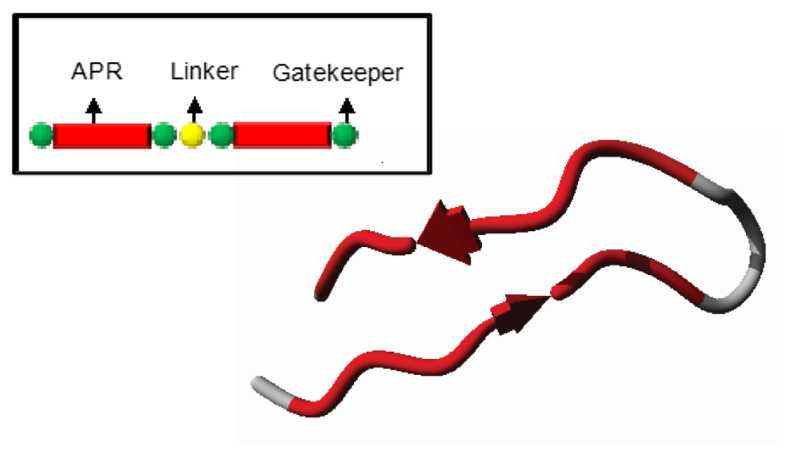
In their most basic design, Pept-ins™ contain tandem repeats of an aggregation prone region flanked by gatekeeper residues and joined by a linker.
The remarkable properties of Pept-ins™ (cellular uptake and efficient biodistribution [5,7,9,10]) make them excellent candidates for therapeutic applications. The Pept-in™ technology allows building drugs based on the primary sequence of the target. They are unique in the sense that Pept-ins™ are active at the protein level yet they can be designed without any information on the structure of the target protein.
The huge potential of the Pept-ins™ technology is apparent when considering its key differentiating features: (1) Designability that allows tuning of the degree of specificity and cross-reactivity (e.g. it is possible to design either broad-spectrum antimicrobial agents or very specific ones). (2) Membrane crossing: The technology allows to target intra-cellular proteins as the peptides can cross the cell/plasma membrane, even in Gram negative strains where this is notoriously difficult. (3) New mode of action: the protein function is inhibited by inducing aggregation during protein production and/or protein processing. This new mode of action opens a completely novel target space for therapeutics development because virtually any protein can be targeted, regardless of its function or cellular location.
The Pept-ins™ technology turns the conventional aim of protein aggregation research (preventing it) on its head and rather uses aggregation to our advantage. By flipping the script like this, profs. Rousseau and Schymkowitz created a unique and entirely novel method to design therapeutics.
Moreover, the selectivity of Pept-ins™ makes it possible to use them for detecting proteins in complex biological mixtures, e.g. detecting biomarkers like C-reactive protein in human blood serum or prostate specific antigen in seminal plasma [11].
The technology has been captured in six patents and is currently licensed to the VIB-KU Leuven spin-off Aelin Therapeutics (Leuven, Belgium). The company is developing the platform technology as a novel human therapeutic modality along antimicrobial and oncological therapeutic programs.
References
- Fernandez-Escamilla AM, Rousseau F, Schymkowitz J, Serrano L (2004) Prediction of sequence-dependent and mutational effects on the aggregation of peptides and proteins. Nature Biotechnology 22 (10):1302-1306. doi:10.1038/nbt1012
- Maurer-Stroh S, Debulpaep M, Kuemmerer N, de la Paz ML, Martins IC, Reumers J, Morris KL, Copland A, Serpell L, Serrano L, Schymkowitz JWH, Rousseau F (2010) Exploring the sequence determinants of amyloid structure using position-specific scoring matrices. Nature Methods 7 (3):237-U109. doi:10.1038/Nmeth.1432
- Betti C, Vanhoutte I, Coutuer S, De Rycke R, Mishev K, Vuylsteke M, Aesaert S, Rombaut D, Gallardo R, De Smet F, Xu J, Van Lijsebettens M, Van Breusegem F, Inze D, Rousseau F, Schymkowitz J, Russinova E (2016) Sequence-Specific Protein Aggregation Generates Defined Protein Knockdowns in Plants. Plant physiology 171 (2):773-787. doi:10.1104/pp.16.00335
- Betti C, Schymkowitz J, Rousseau F, Russinova E (2018) Selective Knockdowns in Maize by Sequence-Specific Protein Aggregation. In: Lagrimini LM (ed) Maize: Methods and Protocols, vol 1676. Methods in Molecular Biology. Humana Press Inc, Totowa, pp 109-127. doi:10.1007/978-1-4939-7315-6_6
- Gallardo R, Ramakers M, De Smet F, Claes F, Khodaparast L, Khodaparast L, Couceiro JR, Langenberg T, Siemons M, Nystrom S, Young LJ, Laine RF, Young L, Radaelli E, Benilova I, Kumar M, Staes A, Desager M, Beerens M, Vandervoort P, Luttun A, Gevaert K, Bormans G, Dewerchin M, Van Eldere J, Carmeliet P, Vande Velde G, Verfaillie C, Kaminski CF, De Strooper B, Hammarstrom P, Nilsson KP, Serpell L, Schymkowitz J, Rousseau F (2016) De novo design of a biologically active amyloid. Science 354 (6313):aah4949-4941 - aah4949-4949. doi:10.1126/science.aah4949
- Bednarska NG, van Eldere J, Gallardo R, Ganesan A, Ramakers M, Vogel I, Baatsen P, Staes A, Goethals M, Hammarstrom P, Nilsson KP, Gevaert K, Schymkowitz J, Rousseau F (2016) Protein aggregation as an antibiotic design strategy. Mol Microbiol 99 (5):849-865. doi:10.1111/mmi.13269
- Khodaparast L, Khodaparast L, Gallardo R, Louros NN, Michiels E, Ramakrishnan R, Ramakers M, Claes F, Young L, Shahrooei M, Wilkinson H, Desager M, Mengistu Tadesse W, Nilsson KPR, Hammarstrom P, Aertsen A, Carpentier S, Van Eldere J, Rousseau F, Schymkowitz J (2018) Aggregating sequences that occur in many proteins constitute weak spots of bacterial proteostasis. Nat Commun 9 (1):866. doi:10.1038/s41467-018-03131-0
- Michiels E, Roose K, Gallardo R, Khodaparast L, Khodaparast L, van der Kant R, Siemons M, Houben B, Ramakers M, Wilkinson H, Guerreiro P, Louros N, Kaptein SJF, Ibanez LI, Smet A, Baatsen P, Liu S, Vorberg I, Bormans G, Neyts J, Saelens X, Rousseau F, Schymkowitz J (2020) Reverse engineering synthetic antiviral amyloids. Nat Commun 11 (1):2832. doi:10.1038/s41467-020-16721-8
- Couceiro JR, Gallardo R, De Smet F, De Baets G, Baatsen P, Annaert W, Roose K, Saelens X, Schymkowitz J, Rousseau F (2015) Sequence-dependent internalization of aggregating peptides. J Biol Chem 290 (1):242-258. doi:10.1074/jbc.M114.586636
- Bednarska NG, van Eldere J, Gallardo R, Ganesan A, Ramakers M, Vogel I, Baatsen P, Staes A, Goethals M, Hammarstrom P, Nilsson KP, Gevaert K, Schymkowitz J, Rousseau F (2015) Protein aggregation as an antibiotic design strategy. Mol Microbiol. doi:10.1111/mmi.13269
- Ganesan A, Debulpaep M, Wilkinson H, Van Durme J, De Baets G, Jonckheere W, Ramakers M, Ivarsson Y, Zimmermann P, Van Eldere J, Schymkowitz J, Rousseau F (2015) Selectivity of aggregation-determining interactions. J Mol Biol 427 (2):236-247. doi:10.1016/j.jmb.2014.09.027